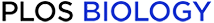Prediction of metal-binding residues and conservation of the spatial arrangement of putative zinc-binding histidines in ZepA homologs.
(A) Plots correspond to the output of the MIB2 metal binding prediction server. The y axis indicates the MIB2 score and the x axis indicates the amino acid positions. The species and the phylum of each ZepA homolog is indicated. Histidines predicted to form the zinc-binding pocket are enclosed in a frame and indicated with a purple bar. Notice that in Gloeothece verrucosa, only 2 of the 3 histidine residues are conserved. The consensus sequence of the putative zinc-binding pocket is shown at the top with histidines residues in purple. The data underlying this figure can be found in S1 Data. (B) Pictures show a close view of the zinc-binding pocket of each ZepA homolog. The structure of each protein was modeled with AlphaFold2. Putative zinc-binding histidine residues are depicted in purple color.
(PPTX)