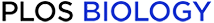The geometrical and topological features of the canonical Rossmann β2-Asp/Glu motif.
(A) Representative carboxylate-ribose bidentate interactions in Rossmann-fold enzymes. Structures were superpositioned by the ribose moiety of their cofactors. One complete backbone is shown (in cartoon), whilst for others, shown are the bound cofactor, the second β-strand (β2), and the interacting Glu or Asp. PDB (Protein Data Bank) IDs and corresponding cofactors: 1JG2, ADN; 3GVI, ADP; 2HMU, ATP; 2XXB, AMP; 1BWC, FAD; 1V5E, FAD; 1EG2, MTA; 2A14, 2PBF, 2AVD (complete structure), SAM; 2GR2, FAD; 1AHH, NAD; 1GEG, NAD; 1GZ6, NAI. (B) The distribution of the interaction angle (α) in structures of proteins with a ribose bound to an Asp/Glu via a bidentate interaction. α is defined by two vectors: v1, going through the CH2-COO- carbons of the interacting Asp/Glu side chain, and v2, going through the C1-C2 carbons of the ribose ring. Gray bars represent the angles in all Rossmann structures with the canonical motif (n = 263). Black bars represent the angles of all the noncanonical bidentate interactions found in both Rossmann and non-Rossmanns enzymes. The PDB Rossmann structures with canonical and noncanonical interaction and their α angles are listed in S1 Data. (C) Representative noncanonical bidentate interactions in non-Rossmann enzymes. PDBs and the corresponding cofactor: 1HO5, ADN; 2J9L (complete structure; the helix carrying the ribose-binding Asp727 is highlighted), ATP; 3S2U, UD1; 1K9Y, AMP; 2ATV, GDP; 1SIW, GDP; 3TE5, NAI; 1I7L, ATP; 4B45, GSP. These structures are shown individually in S6 Fig.