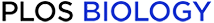Conservation of amino acid sequences between Hv1 orthologues.
(A) Multiple sequence alignment of animal and coccolithophore Hv1 proteins. The multiple sequence alignment indicates transmembrane residues conserved between animal and coccolithophore Hv1 proteins. Shading indicates residues that are identical or similar in 80% of the sequences (BLOSUM62 matrix), and boxes represent predicted transmembrane domains in human Hv1. The three arginine residues in S4 required for voltage sensing are conserved (numbers below alignment correspond to R205, R208, and R211 in human Hv1). Many of the acidic residues (aspartate and glutamate) appear conserved (D112, E119, D123, E153, D174), although Asp185 is replaced by glutamate and Glu171 is not conserved. Several of these acidic residues are conserved amongst voltage sensor domain proteins and may function in voltage sensing [29]. In contrast, the other basic residues (lysine) are not conserved in coccolithophores (K125, K157, K169, K221). Polar serine residues (S143, S181) are conserved in E. huxleyi, although Ser181 is replaced by alanine in C. pelagicus. Similar sequences identified in the diatoms P. tricornutum and T. pseudonana are also shown in the alignment. The arrows indicate external histidine residues required for Zn2+ inhibition in human Hv1 [19] and external histidine residues which are conserved in algal Hv1 proteins (the numbers above the alignment indicate their position in EhHv1). (B) Multiple sequence alignment indicating conserved domains in transmembrane regions of voltage sensor proteins. Hv1 proteins from animals (Hs, Homo sapiens; Dr, Danio rerio; Ci, Ciona intestinalis) and coccolithophores (Eh, Emiliania huxleyi; Cp, Coccolithus pelagicus) are aligned to transmembrane regions S1–S4 from K+ channels from humans, Drosophila, plants, and prokaryotes and also to the prokaryote Na+ channel NaChBac. Shading indicates identical residues in 5 out of 10 sequences displayed. In this analysis the only conserved residues that are exclusive to Hv1 proteins correspond to human Hv1 residues D112, D123, and S143. (C) Phylogenetic analysis of known Hv1 proteins. The tree was generated by a maximum likelihood analysis of available Hv1 protein sequences. 100 bootstraps were performed.