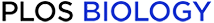Pharmacogenetic activation of the hM3Dq-expressing rat rostromedial tegmental nucleus (RMTg) neurons.
Schematic representation of the Cre-independent adeno-associated virus (AAV) vectors expressing hM3Dq receptors under the control of hSyn promoter. hSyn, human synapsin promoter; WPRE, woodchuck hepatitis virus post-transcriptional regulatory element; CNO, clozapine-N-oxide; IP, intraperitoneal. (B) The coronal section shows the superimposed virus-injected area in 9 rats numbered with the same color characters as the closed curves. The bilateral shaded areas indicate the rat RMTg locations. (C) mCherry immunolabeling (red) indicates hM3Dq-expressing neurons of the RMTg. IPN, interpeduncular nucleus; ml, medial lemniscus. (D) Representative photomicrographs of the RMTg depicting mCherry expression (red), γ-amino butyric acid (GABA) immunoreactivity (green), 4,6-diamidino-2-phenylindole (DAPI, blue), and merge (yellow) images from a rat microinjected with Cre-independent AAV vectors containing hM3Dq. (E, F) CNO (E), not saline (F), drives c-Fos expression in hM3Dq-expressing neurons in the RMTg. (G) Quantification of c-Fos staining in mCherry-expressing RMTg neurons after saline or CNO injection in rats. **p < 0.01 versus saline by unpaired t tests (n = 3, per group). (H) A typical example of voltage trace recorded from an hM3Dq+ rat RMTg neuron during application of CNO in a brain slice. CNO application (horizontal bar) increased the action potential firing. (I) Merged image of mCherry and biocytin showing a fluorescent view of the patched cell. Left panel: biocytin-labeled neuron; right panel: cell with green (biocytin; top), red (hM3Dq-mCherry; middle), and yellow (merged; bottom) fluorescence. (J, K) Average membrane potential of RMTg hM3Dq+ neurons (J, n = 7 cells), but not hM3Dq− neurons (K, n = 5 cells), was significantly increased by CNO application. The results from each cell are shown on the scatterplot. **p < 0.01 versus saline by paired t tests. Underlying data can be found in S1 Data.